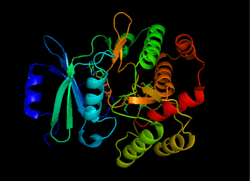
Kanamycin kinase
| kanamycin kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
|
Structure of APH(3'), taken from 1L8T[1] | |||||||||
| Identifiers | |||||||||
| EC number | 2.7.1.95 | ||||||||
| CAS number | 62213-36-9 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
Aminoglycoside-3'-phosphotransferase (APH(3')), also known as aminoglycoside kinase, is an enzyme that primarily catalyzes the addition of phosphate from ATP to the 3'-hydroxyl group of a 4,6-disubstituted aminoglycoside, such as kanamycin.[2] However, APH(3') has also been found to phosphorylate at the 5'-hydroxyl group in 4,5-disubstituted aminoglycosides, which lack a 3'-hydroxyl group, and to diphosphorylate hydroxyl groups in aminoglycosides that have both 3'- and 5'-hydroxyl groups.[2][3] Primarily positively charged at biological conditions, aminoglycosides bind to the negatively charged backbone of nucleic acids to disrupt protein synthesis, effectively inhibiting bacterial cell growth.[4] APH(3') mediated phosphorylation of aminoglycosides effectively disrupts their mechanism of action, introducing a phosphate group that reduces their binding affinity due to steric hindrances and unfavorable electrostatic interactions.[5] APH(3') is primarily found in certain species of gram-positive bacteria.[6][7][8]

This enzyme belongs to the family of transferases, specifically those transferring phosphorus-containing groups (phosphotransferases) with an alcohol group as acceptor. The systematic name of this enzyme class is ATP:kanamycin 3'-O-phosphotransferase. This enzyme is also called neomycin-kanamycin phosphotransferase.[9]
Structure
APH(3’) thermodynamically favors a dimer form of two identical APH(3’) monomers that are connected by two disulfide bonds between Cys19 and Cys156, with the active sites facing each other.[2][10] However, the large distance between the two monomers’ active sites suggests that they are independent of each other, and do not operate in a cooperative fashion. Additionally, dimerization of APH(3’) does not affect the activity of the enzyme.[2][10][11]
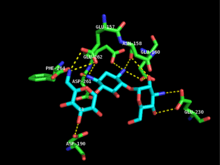
Each monomer consists of two lobes, the beta-sheet rich N-terminus and alpha-helix rich C-terminus, with a twelve amino acid region connecting the two. The N-terminal lobe is composed of 5 antiparallel ß-sheets, with an α-helix between sheets 3 and 4. The C-terminal lobe is divided into a central core region (two α-helices and a hairpin-loop followed by four ß-sheets), an insert region (two α-helices connected by a loop structure), and a C-terminal region (two α-helices).[11] The resulting pocket that is encapsulated by the two lobes make up the enzyme active site.[2] This pocket is largely composed of negatively charged amino acid residues, which stabilize the positive charge of and orient the substrate in the active site. Additionally, this pocket is thought to contribute to the promiscuity of the enzyme, allowing it to take in and stabilize several different kinds of aminoglycosides.[6]
Mechanism
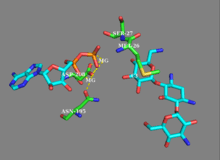
While earlier studies of APH(3') supported a mechanism involving the nucleophilic attack of γ-phosphate by the 3'-hydroxyl, more recent studies suggest that APH(3’) catalyzes the transfer of the γ-phosphate from ATP to an aminoglyocside through a dissociative mechanism, where deprotonation of the substrate is not critical to phosphate transfer, but instead the stabilization of a metaphosphate transition state.[8][12] Additionally, APH(3’) has a nucleotide positioning loop (NPL) that closes down on the enzyme active site after binding ATP, facilitating the phosphorylation of the 3’-hydroxyl group. Key to correctly positioning the phosphate group are Ser27 and Met26 residues. Initially, two magnesium ions stabilized by Asn195 and Asp208 facilitate the binding of ATP in the active site and orient the ß- and γ-phosphate groups. The NPL then undergoes a conformational change to form a hydrogen bond between Ser27 and the ß-phosphate group. Upon binding of substrate, APH(3') undergoes another conformational change to orient Ser27 such that its amide backbone disrupts the alignment of ß-phosphate and γ-phosphate, weakening the γ-phosphate bond. The amide backbone of Met26 forms a hydrogen bond with the metaphosphate to stabilize the transition state, as a magnesium ion (designated Mg1) then lengthens the γ-phosphate bond, breaking it and effectively phosphorylating the hydroxyl group.[8]

Evolution and Biological Function
The central core region of APH(3’) has a high degree of conformational similarity to regions of serine/tyrosine and threonine protein kinases, functionally equivalent enzymes found in eukaryotes. Additionally, X-ray crystallography and mutagenesis of key active site residues supports claims that APH(3’) and eukaryotic protein kinases are related, despite sharing less than 10% of total residue content.[2][8][11] Several studies have suggested that serine/tyrosine/threonine protein kinases, once thought to only occur in eukaryotes, are also found in the prokaryotes.[13][14] Additionally, it has been found that aminoglycoside biosynthesis requires phosphorylation of hydroxyl groups during certain steps of synthesis. Thus, it has been speculated that APH(3’) and other protein kinases originate from enzymes from the metabolic pathway for aminoglycosides, and developed in order to counteract the toxic effects of these antibiotics in the host bacterial cell.[11][15]
Use in Research
Aminoglycoside resistance genes are commonly used in the realm of genetic engineering in order to select for correctly transformed bacterial organisms. When constructing a vector plasmid, including antibiotic resistance in the vector is crucial to effectively expressing the gene of interest. Antibiotics, such as the aminoglycosides kanamycin or neomycin, are added to the cultures during growth phases in order to selectively destroy the cells that did not effectively take up the plasmid.
References
- ↑ Fong, DH, Berghuis, AM (2002). "Crystal Structure Of 3',5"-Aminoglycoside Phosphotransferase Type IIIa ADP Kanamycin A Complex". doi:10.2210/pdb1l8t/pdb.
- 1 2 3 4 5 6 7 Wright, GD, Thompson, PR (1999). "Aminoglycoside phosphotransferases: proteins, structure, and mechanism.". Front Biosci. 4: D9–21. doi:10.2741/wright. PMID 9872733.
- ↑ Thompson, PR, Hughes, DW, Wright, GD (1996). "Regiospecificity of aminoglycoside phosphotransferase from Enterococci and Staphylococci (APH(3')-IIIa).". Biochemistry. 35: 8686–95. doi:10.1021/bi960389w. PMID 8679631.
- ↑ Cavallo, G, Martinetto, P (1981). "The mechanism of action of aminoglycosides". G Batteriol Virol Immunol. 74: 335–46. PMID 6182050.
- ↑ Kotra, LP, Haddad J, Mobashery, S (2000). "Aminoglycosides: Perspectives on Mechanisms of Action and Resistance and Strategies to Counter Resistance". Antimicrobial Agents and Chemotherapy. 44 (12): 3249–56. doi:10.1128/aac.44.12.3249-3256.2000. PMC 90188
 . PMID 11083623.
. PMID 11083623. - 1 2 Fong, DH, Berghuis, AM (2002). "Substrate promiscuity of an aminoglycoside antibiotic resistance enzyme via target mimicry.". The EMBO Journal. 15 (10): 2323–31. doi:10.1093/emboj/21.10.2323. PMC 126009
 . PMID 12006485.
. PMID 12006485. - ↑ Gray, GS, Fitch WM (1983). "Evolution of antibiotic resistance genes: the DNA sequence of a kanamycin resistance gene from Staphylococcus aureus.". Mol Bio Evol. 1 (1): 57–66. PMID 6100986.
- 1 2 3 4 5 Thompson, PR, Boehr, DD, Berghuis, AM, Wright, GD (2002). "Mechanism of Aminoglycoside Antibiotic Kinase APH(3')-IIIa: Role of the Nucleotide Positioning Loop". Biochemistry. 41 (22): 7001–7. doi:10.1021/bi0256680. PMID 12033933.
- ↑ McKay GA, Wright GD (1996). "Catalytic mechanism of enterococcal kanamycin kinase (APH(3')-IIIa): viscosity, thio, and solvent isotope effects support a Theorell-Chance mechanism". Biochemistry. 35 (26): 8680–5. doi:10.1021/bi9603884. PMID 8679630.
- 1 2 McKay, GA, Thompson, PR, Wright, GD (1994). "Broad spectrum aminoglycoside phosphotransferase type III from Enterococcus: overexpression, purification, and substrate specificity.". Biochemistry. 33 (22): 6936–44. doi:10.1021/bi00188a024. PMID 8204627.
- 1 2 3 4 Hon, WC, McKay, GA, Thompson, PR, Sweet, RM, Yang, DSC, Wright, GD, Berhuis, AM (1997). "Structure of an Enzyme Required for Aminoglycoside Antibiotic Resistance Reveals Homology to Eukaryotic Protein Kinases.". Cell. 89 (6): 887–95. doi:10.1016/s0092-8674(00)80274-3. PMID 9200607.
- ↑ Boehr, DD, Thompson, PR, Wright, GD (2001). "Molecular mechanism of aminoglycoside antibiotic kinase APH(3')-IIIa: roles of conserved active site residues.". J Biol Chem. 276 (26): 23929–36. doi:10.1074/jbc.m100540200. PMID 11279088.
- ↑ Kennelly, PJ (1996). "Fancy meeting you here! A fresh look at "prokaryotic" protein phosphorylation.". J Bacteriol. 178 (16): 4759–64. PMID 8759835.
- ↑ Zhang, CC (1996). "Bacterial signalling involving eukaryotic-type protein kinases.". Mol Microbiol. 20 (1): 9–15. doi:10.1111/j.1365-2958.1996.tb02483.x. PMID 8861199.
- ↑ Pierpersberg, W, Distler, J, Heinzel, P, Perez-Gonzalaez, JA (1988). "Antibiotic resistance by modification: Many resistance genes could be derived from cellular control genes in actinomycetes - a hypothesis". Actinomycetologica. 2 (2): 83–98. doi:10.3209/saj.2_83.
Further reading
- Doi O, Ogura M, Tanaka N, Umezawa H (Sep 1968). "Inactivation of kanamycin, neomycin, and streptomycin by enzymes obtained in cells of Pseudomonas aeruginoa". Applied Microbiology. 16 (9): 1276–81. PMC 547640
 . PMID 4970990.
. PMID 4970990. - Dolin MI (Mar 1957). "The Streptococcus faecalis oxidases for reduced diphosphopyridine nucleotide. III. Isolation and properties of a flavin peroxidase for reduced diphosphopyridine nucleotide". The Journal of Biological Chemistry. 225 (1): 557–73. PMID 13416259.